|
11/18/2023 0 Comments Ires sequences Proteins were synthesized in reaction mixtures containing methionine, DNVR2 RNA, and DNVR19 RNA (lane 1) or DNVR22 RNA (lane 2). (B) Cotranslation of DNVR2 helper mRNA with PV template RNAs. DNVR22 RNA possesses the 5′-terminal 123 nt (cloverleaf) and the 3′ NTR-poly(A) tail of PV. Negative-strand RNA synthesis with a minimally sufficient RNA template. The mobilities of DNVR10 and DNVR17 negative-strand RNAs are indicated. Radiolabeled negative-strand RNAs were fractionated by electrophoresis in a denaturing methyl mercury hydroxide agarose gel and detected by phosphorimaging. Negative-strand RNA synthesis was assayed in the presence (lanes 1 and 3) and absence (lanes 2 and 4) of 2 mM guanidine HCl (GuHCl). 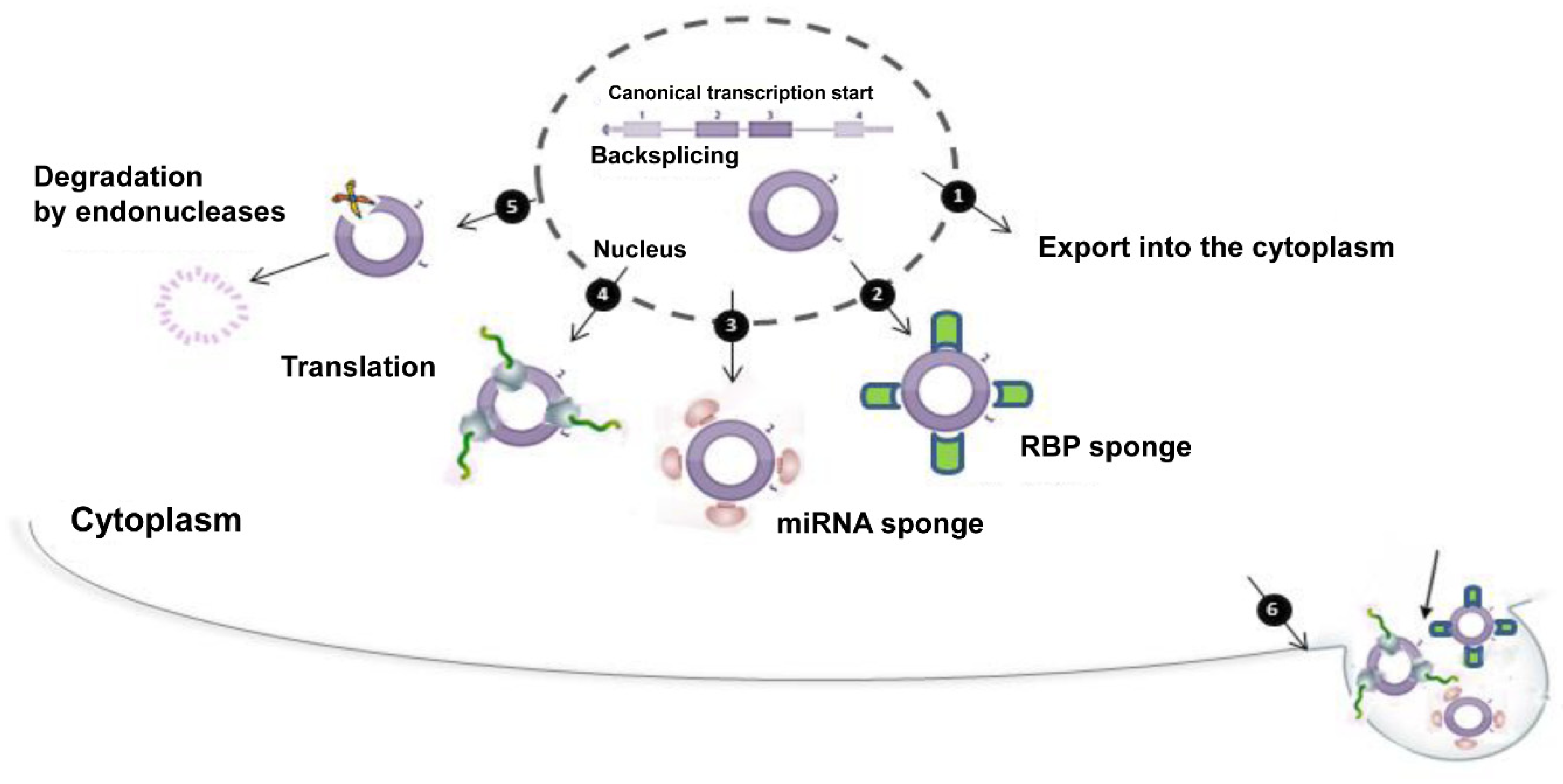
Preinitiation RNA replication complexes contained either DNVR10 RNA templates (lanes 1 and 2) or DNVR17 RNA templates (lanes 3 and 4). Preinitiation RNA replication complexes formed during cotranslation reactions were assayed for the ability to synthesize negative-strand RNA as described in Materials and Methods. Proteins were synthesized in reaction mixtures containing methionine, DNVR2 RNA, and DNVR10 RNA (lane 1) or DNVR17 RNA (lane 2). DNVR17 possesses a deletion of IRES nt 124 to 742. DNVR10 possesses a deletion of IRES nt 220 to 742. Negative-strand RNA synthesis with a template with a complete IRES deletion. Mobilities of DJB14 and DNVR10 negative-strand RNAs are indicated. Radiolabeled negative-strand RNAs fractionated by electrophoresis in a denaturing methyl mercury hydroxide agarose gel were detected by phosphorimaging. Negative-strand RNA synthesis was assayed in the presence (lanes 1 and 3) and absence (lanes 2 and 4) of 2 mM guanidine HCl. Preinitiation RNA replication complexes contained either DJB14 (lanes 1 and 2) or DNVR10 (lanes 3 and 4) RNA templates. 
Proteins were synthesized in reaction mixtures containing methionine, DNVR2 RNA, and DJB14 RNA (lane 1) or DNVR10 RNA (lane 2). (C) Cotranslation of DNVR2 helper mRNA with DJB14 and DNVR10 PV template RNAs. DNVR10 RNA possesses a deletion of IRES nt 220 to 742. DJB14 RNA possesses a wild-type PV IRES, a small ORF encoding a COOH-terminal fragment of 3D Pol (Δ3D Pol), and the PV 3′ NTR-poly(A) tail. This mRNA possesses the 5′ NTR of HCV, an ORF encoding the PV replication proteins, and the 3′ NTR-poly(A) tail of PV. IRES stem-loops IV, V, and VI are dispensable for negative-strand RNA synthesis. These results further suggest that poliovirus RNA replication is not absolutely dependent on any protein-RNA interactions involving the IRES. These results suggest that RNA replication is not directly dependent on a template RNA first functioning as an mRNA. Intriguingly, although small viral RNAs lacking the IRES replicated efficiently, the replication of genome length viral RNAs was stimulated by the presence of the IRES. The IRES was dispensable for both negative- and positive-strand RNA syntheses. A series of deletions revealed that no RNA elements of either the viral open reading frame or the IRES were required in cis for negative-strand RNA synthesis. Genetic complementation with helper mRNAs allowed us to create preinitiation RNA replication complexes containing RNA templates with defined deletions in the viral open reading frame and the IRES. 
To evaluate the role of the IRES in poliovirus RNA replication, we exploited the advantages of cell-free translation-replication reactions and preinitiation RNA replication complexes. The poliovirus internal ribosome entry site (IRES) in the 5' nontranslated region (NTR) has been implicated as a cis-active RNA required for both viral mRNA translation and viral RNA replication. Cis-acting RNA sequences and structures in the 5' and 3' nontranslated regions of poliovirus RNA interact with host translation machinery and viral replication proteins to coordinately regulate the sequential translation and replication of poliovirus RNA. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
